SeaView
5.0.4Advanced and portable program for multiple sequence alignment and molecular phylogeny analysis that reads and writes various files, such as NEXUS, MSF, CLUSTAL, FASTA, PHYLIP, MASE and Newick
SeaView is a software application designed specifically for helping you study and analyze sequence alignments as well as molecular phylogeny. It integrates abilities for reading and writing various file formats, such as NEXUS, MSF, CLUSTAL, FASTA, PHYLIP, MASE and Newick files.
This is a standalone program that can be run directly on your computer without being installed in a way that alters your Windows registry configuration information. You can store the utility on any storage devices such as USB flash drives.
The best part when you are using portable tools is that you don’t need administrator privileges to run the programs as they don’t leave files or settings on the host computer.
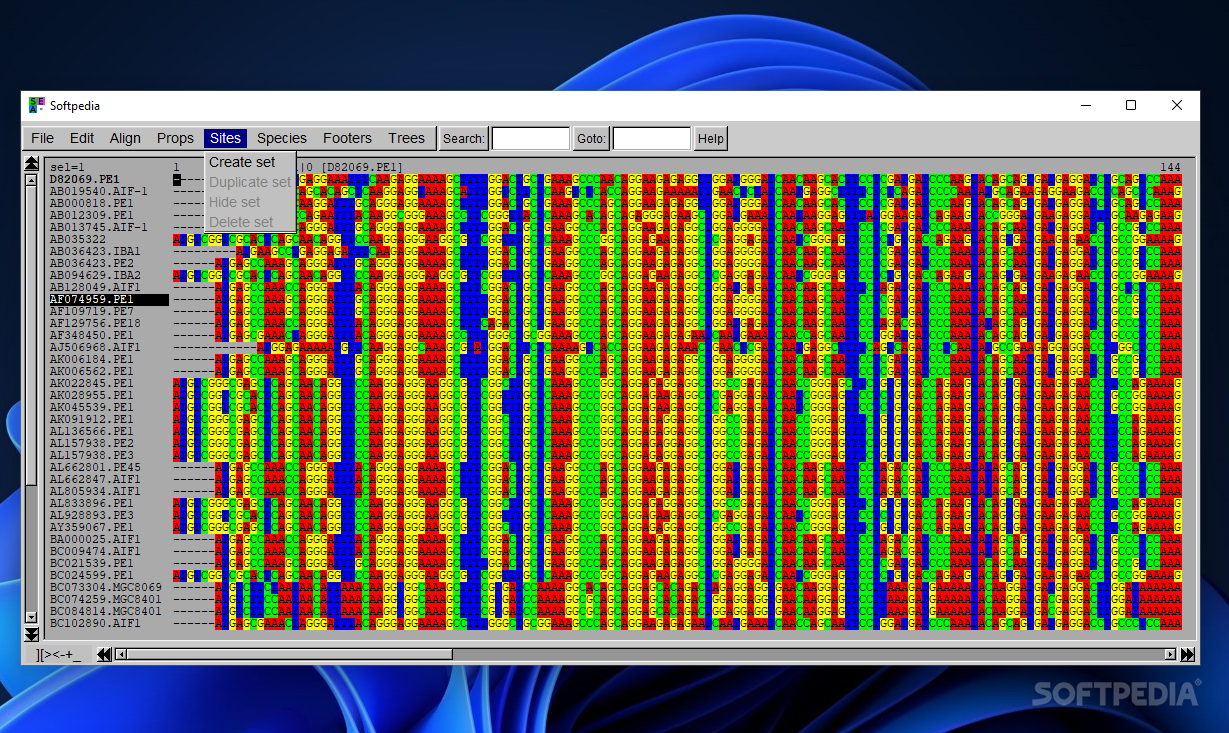
Although the GUI doesn’t not impress much in the visual department, it is actually practical and easy to work with. Drag-and-drop support is also on the feature list so you may quickly import data from an external file.
You can make use of keyboard shortcuts for exploring data or press mouse clicks for selecting items and moving selected sequences to another place in an alignment. The tool can also be run using the command-line.
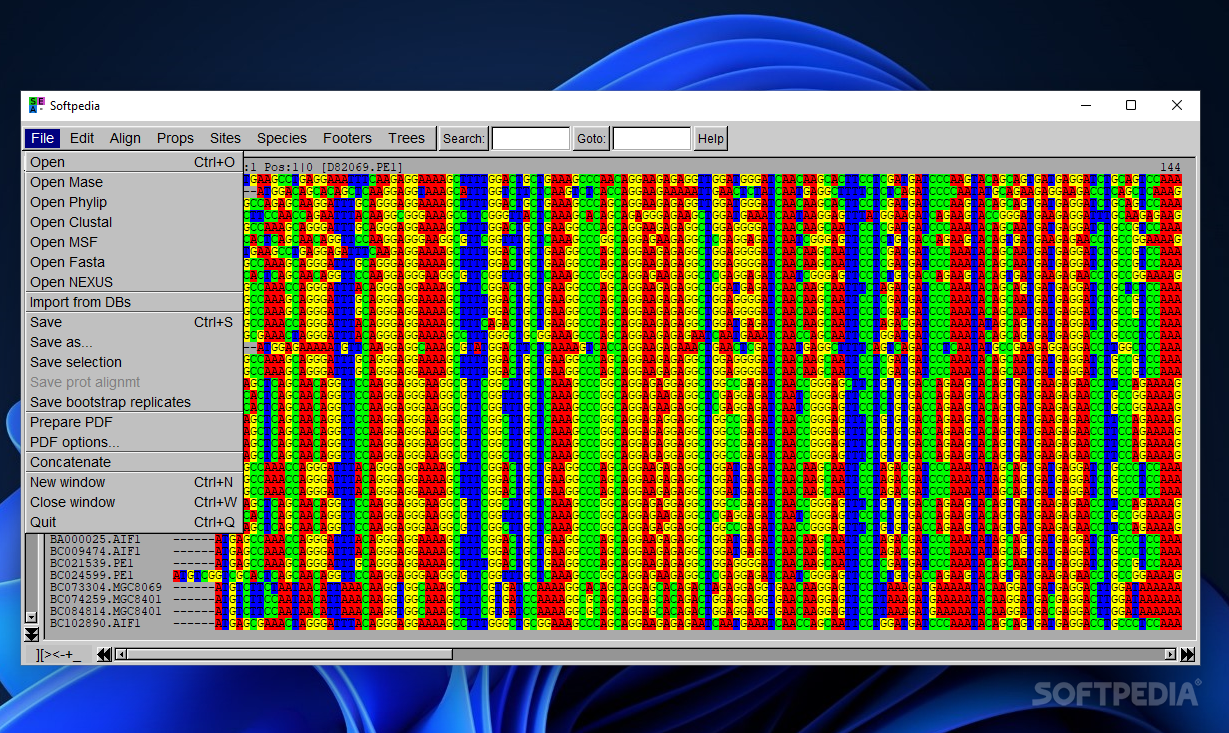
Aside from the supported file formats that have been enumerated in the introduction, you can add information from various databases, such as EMBL, GenBank and SwissProt/UniProt.
Additionally, you can save the alignment in the current file or export data to a file, save the alignment to PDF file format (you are given the freedom to alter the font, block and paper size, change the color, and output only variable sites of the alignment and their positions). You are allowed to add one alignment to the end of another one by name or rank.
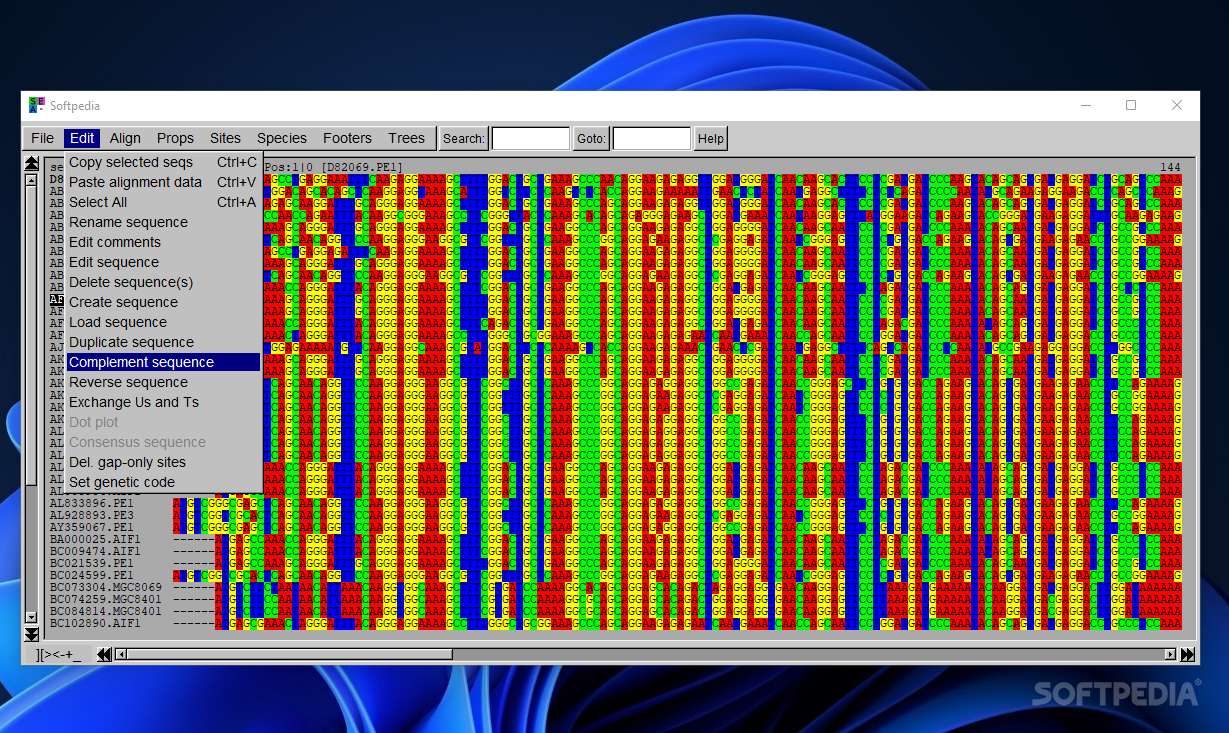
SeaView gives you the possibility to copy the selected sequences to the clipboard, paste alignment data from the clipboard, select all sequences from the alignment, rename the currently selected sequence, edit comments, as well as alter a sequence by pasting external data or opening two editing sequence panels and transferring info between them.
Other handy tasks that can be applied to sequences refer to deleting, creating and loading them. Plus, you can duplicate and reverse them, perform a dot plot analysis, delete all gap sites from the alignment and set the genetic code for translating to protein the selected sequence(s).
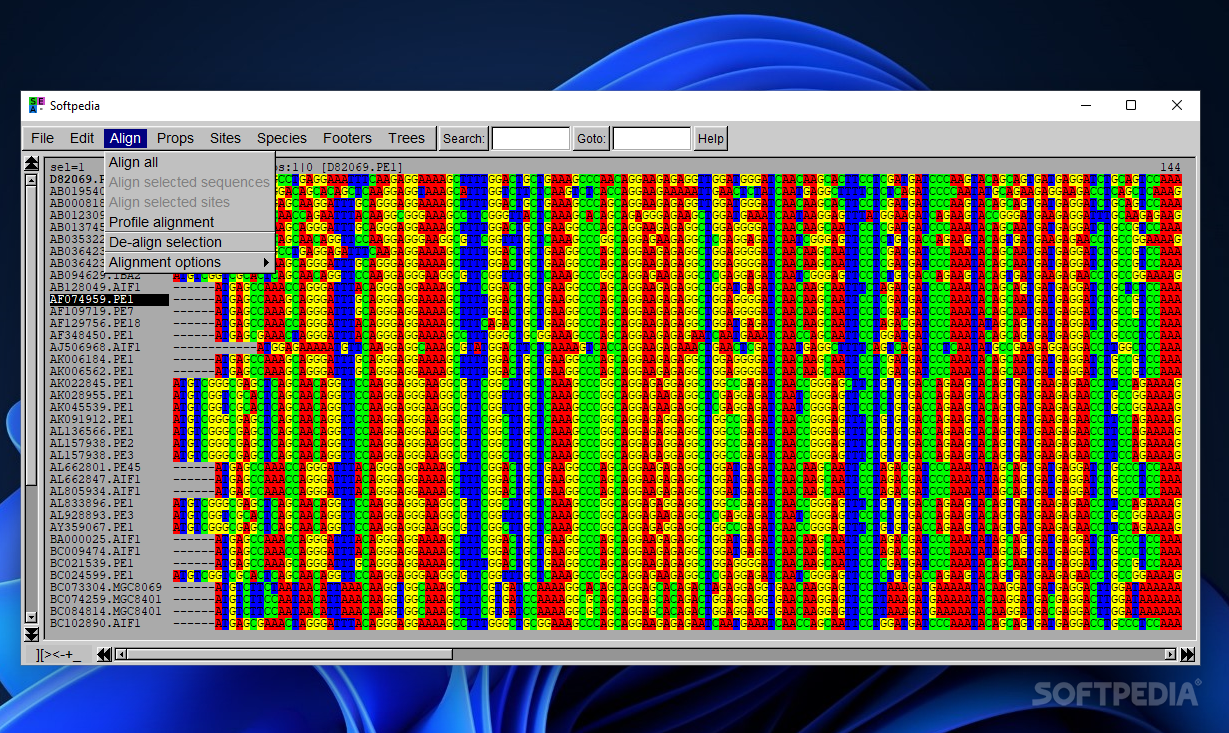
When it comes to actions that help you manage alignments, you may run the target alignment on all or selected sequences, selected sites or a profile (more specifically a group of pre-aligned sequences).
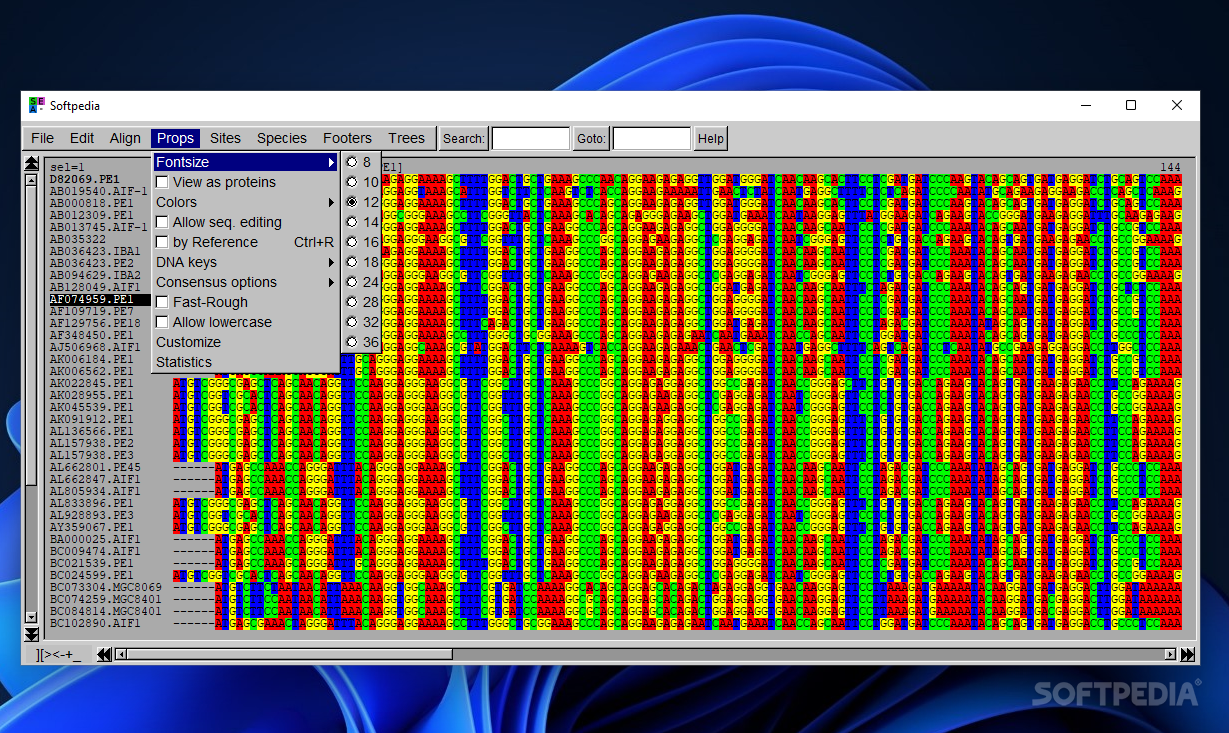
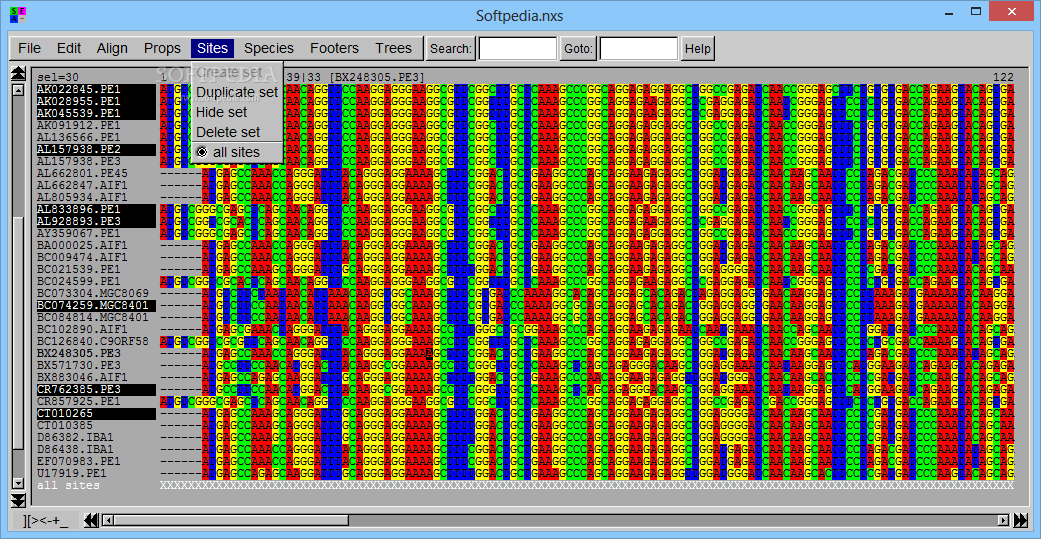
SeaView lets you set the font size used to display sequences, display DNA sequences as protein ones, use color rules for defining the protein sequences, generate statistics about base or amino acid sequence compositions, transition and transversion counts, specify the parts of a multiple alignment to be stored for further analysis, create and store species sets if the Mase or NEXUS format is used, insert comment lines at the bottom of the screen, and carry out searches.
Furthermore, you may compute, draw, import and save DNA or protein phylogenetic trees, plot, save print and copy phylogenetic trees, and perform a ‘dot plot’ analysis of two sequences.
All in all, SeaView proves to be an advanced program for multiple sequence alignment and molecular phylogeny, and is suitable especially for power users.
Portable package
This is a standalone program that can be run directly on your computer without being installed in a way that alters your Windows registry configuration information. You can store the utility on any storage devices such as USB flash drives.
The best part when you are using portable tools is that you don’t need administrator privileges to run the programs as they don’t leave files or settings on the host computer.
User interface
Although the GUI doesn’t not impress much in the visual department, it is actually practical and easy to work with. Drag-and-drop support is also on the feature list so you may quickly import data from an external file.
You can make use of keyboard shortcuts for exploring data or press mouse clicks for selecting items and moving selected sequences to another place in an alignment. The tool can also be run using the command-line.
Importing/exporting options
Aside from the supported file formats that have been enumerated in the introduction, you can add information from various databases, such as EMBL, GenBank and SwissProt/UniProt.
Additionally, you can save the alignment in the current file or export data to a file, save the alignment to PDF file format (you are given the freedom to alter the font, block and paper size, change the color, and output only variable sites of the alignment and their positions). You are allowed to add one alignment to the end of another one by name or rank.
Editing features and alignment options
SeaView gives you the possibility to copy the selected sequences to the clipboard, paste alignment data from the clipboard, select all sequences from the alignment, rename the currently selected sequence, edit comments, as well as alter a sequence by pasting external data or opening two editing sequence panels and transferring info between them.
Other handy tasks that can be applied to sequences refer to deleting, creating and loading them. Plus, you can duplicate and reverse them, perform a dot plot analysis, delete all gap sites from the alignment and set the genetic code for translating to protein the selected sequence(s).
When it comes to actions that help you manage alignments, you may run the target alignment on all or selected sequences, selected sites or a profile (more specifically a group of pre-aligned sequences).
Configuration settings and other powerful tools
SeaView lets you set the font size used to display sequences, display DNA sequences as protein ones, use color rules for defining the protein sequences, generate statistics about base or amino acid sequence compositions, transition and transversion counts, specify the parts of a multiple alignment to be stored for further analysis, create and store species sets if the Mase or NEXUS format is used, insert comment lines at the bottom of the screen, and carry out searches.
Furthermore, you may compute, draw, import and save DNA or protein phylogenetic trees, plot, save print and copy phylogenetic trees, and perform a ‘dot plot’ analysis of two sequences.
Bottom line
All in all, SeaView proves to be an advanced program for multiple sequence alignment and molecular phylogeny, and is suitable especially for power users.
7.6 MB
Info
Update Date
Jun 18 2020
Version
5.0.4
License
GPL
Created By
Manolo Gouy
Related software Development